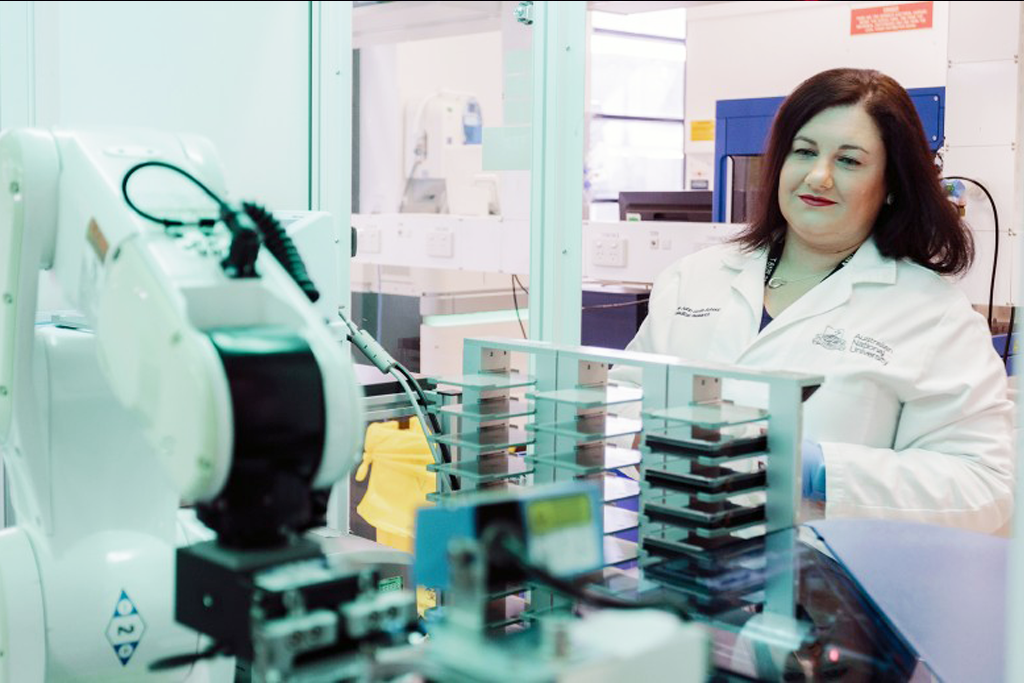
Frequently referred to as the building blocks of life, proteins are vital in all cells in all living organisms.
That puts ribosomes—the structures responsible for protein synthesis—at the heart of a cell’s survival, growth, and proliferation.

“ Ribosomes are dynamic factories which are constantly built and turned over inside cells; this is a process known as ribosome biogenesis,” said Dr Amee George, Phenomics Australia Scientific Leader and Fellow and Lead of the ANU Centre for Therapeutic Discovery (ACTD).
“ To build these factories, we need all of the ‘bricks’ of the ribosome—ribosomal proteins—as well as ribosomal RNA, the ‘mortar’ required to hold these bricks together, to come together.”
A human ribosome, for example, contains approximately 80 different ribosomal proteins and four different ribosomal RNAs. Depending on the cell type, a cell may harbour 10 million or more ribosomes at once.
But not all ribosomes are built correctly.
Deficiencies in the molecular ‘bricks’ and ‘mortar’ required for ribosome biogenesis can disrupt the assembly of ribosomes and cause diseases known as ribosomopathies, including Diamond-Blackfan Anemia (DBA).
“ Individuals diagnosed with a ribosomopathy can present with a variety of different symptoms,” said Dr George, who has a research interest in DBA, “However, one of the key features of many ribosomopathies is bone marrow failure, which can severely impact on blood cell formation.”
Bone marrow failure is characterised by a patient’s inability to make enough healthy blood cells—in the case of DBA, specifically red blood cells, resulting in anemia which requires lifelong clinical management.
For years, Dr George and colleagues have been seeking therapeutic targets to treat bone marrow failure.
“To this end, we needed to understand the mechanism which underpins why mutations in specific ribosomal protein genes lead to bone marrow failure,” she said.
And that set off a research project that has spanned more than eight years.
This story was first published on JCSMR’s website.
When a cell is stressed
A decade ago, scientists investigating the bone marrow of individuals with DBA observed increased levels of a specific protein known as p53.
“p53 is an important tumour suppressor protein in the human body that acts as a cellular ‘handbrake’,” Dr George explained, “If a cell detects a problem, p53 protein then accumulates and triggers a series of molecular events which stops the cell from growing or leads to its self-destruction.”
For example, when a cell is exposed to some types of environmental or chemical stresses, like DNA damage, thermal shock, and, importantly, altered ribosome biogenesis, the cell’s nucleolus—the pivotal site for ribosome biogenesis—detects the stress and initiates a response which allows p53 protein to accumulate in the cell and determine its fate.
If the gene encoding p53 is lost or mutated, it will effectively release this ‘handbrake’ and allow cells to grow uncontrolled, as seen in over 50% of cancers.
Researchers suspect this stress response is the primary reason for bone marrow failure in DBA patients with specific mutated ribosomal protein genes.
“However, the key components underlying this process have not been well established until now,” said Dr George.
Adopting an approach known as functional genomics, her team perturbed gene expression of every individual gene in the genome to identify all genes that play a part in the p53 stress response.
“This approach is extremely powerful as it can be adapted to anything – as long as we have a robust mechanism to measure how the cells respond when depleted of individual genes.”
As an example, for scientists interested in cancer metastasis, this approach can be used to prevent the expression of individual genes in metastatic cancer cells, and identify the genes which are important in preventing metastasis from happening.
This approach helps to provide information on what specific genes or proteins are involved in this process, along with a potential biomarker that could be therapeutically targeted to treat disease.
“But with some 18,000 to 20,000 protein-coding genes located within the human genome, you definitely don’t want to do all that manually,” said Dr George.
Thankfully, they don’t have to.
Using state-of-the-art robotic and liquid handling facilities in two Phenomics Australia nodes in the ACTD and the Victorian Centre for Functional Genomics (VCFG) at the Peter MacCallum Cancer Centre, her team was able to complete these screens in a matter of months.
From this, her team were able to deduce the genes which, in effect, control cellular p53 protein levels.
Intriguingly, ribosome biogenesis-related genes, when disrupted, were the most potent at increasing p53 protein within cells.
In fact, among the 80 human ribosomal protein genes tested in the study, almost 80% of them led to p53 accumulation when perturbed.
Nevertheless, two particular genes, encoding ribosomal proteins RPL5 and RPL11, fell into the minority.
Dual roles of the protein duo
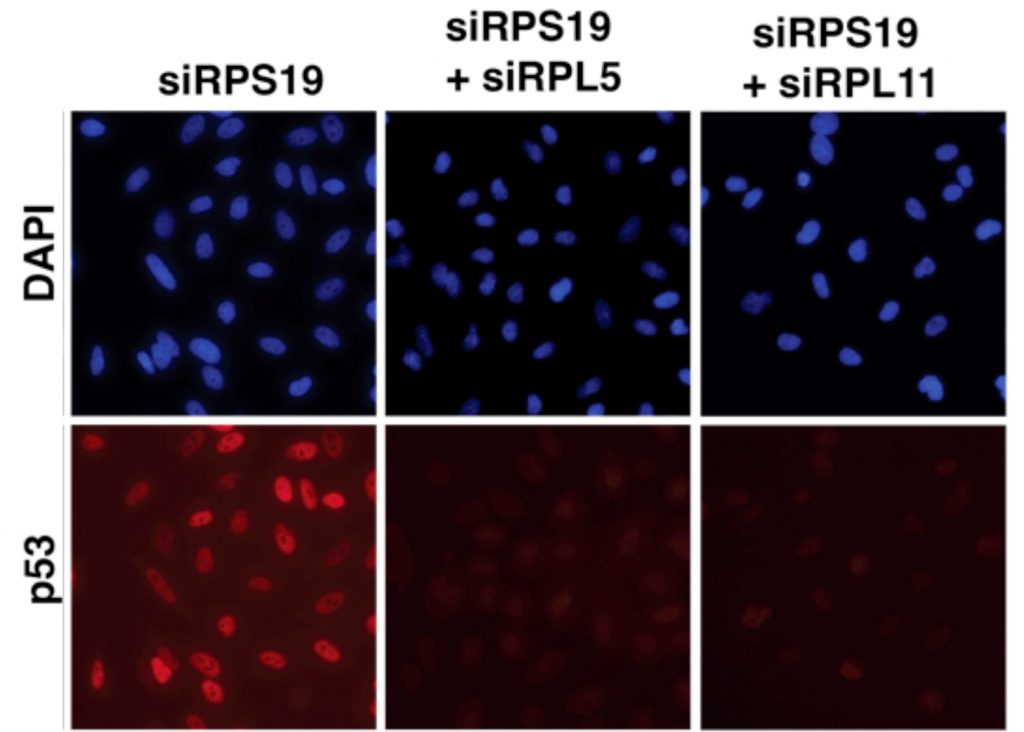
“RPL5 and RPL11 are specifically important because, together, they form a complex that is incorporated into ribosomes just before the start of protein synthesis,” said Dr George.
Under stress conditions, rather than joining the ribosome, the complex instead interacts with a protein called MDM2, which usually regulates p53 activity.
“This means that the presence of the complex and expression of RPL5 and RPL11 are critical for p53 accumulation,” said Dr George.
Further functional unbiased screens revealed that RPL5 and RPL11 are the only ribosomal proteins essential for stabilising p53 in response to disruptions to ribosome biogenesis— confirming they are so important that their depletion terminates the stress.
“Our study provides a detailed picture of key components required for initiating and modulating this stress response in cells,” said Dr George, who recently published the study in Cell Reports*.
Therefore, researchers could find ways to modulate the stress response by deciphering and targeting components relevant to stress response due to ribosomal disruption, such as RPL5 and RPL11.
“This information will be vital for developing novel therapeutic approaches for treating ribosomopathies and other disorders, like some forms of cancer, where this stress mechanism is extremely important,” said Dr George.
Based on their findings, Dr George and collaborators have also conducted drug repurposing screens to seek new uses for therapies designed and approved for treating other diseases to treat DBA.
The team has now tested approximately 3500 FDA-approved drugs. In addition to this, Dr George also has an interest in developing novel drugs targeting this process.
“What we have ascertained from our studies thus far is a deeper understanding of the global pathways involved in the stress response elicited by the nucleolus and potential targets we can go after therapeutically to treat bone marrow failure in individuals with ribosomopathies.”
Particularly for individuals with DBA, therapeutic options for treatment remain limited and are highly invasive. Breakthroughs in understanding the molecular mechanisms underlying the disease may fuel life-changing translational discoveries.
“ Medical researchers focus on making discoveries which can be translated into advances that will make a real-world difference to those living with medical conditions,” said Dr George.

“But to make significant progress on some of these particular diseases and disorders, we need to first understand the molecular basis underlying them.”
This work is dedicated to the memory of Associate Professor Kate Hannan, who was the first author of the published study. The research was funded by the NHMRC and Captain Courageous Foundation (now part of Maddie Riewoldt’s Vision).
The ACTD, located within the John Curtin School of Medical Research, is funded by the ANU, Australian Cancer Research Foundation (ACRF) and ACT Health, and also supported by the NCRIS funding bodies Phenomics Australia and Therapeutic Innovation Australia. The ACTD is a dedicated facility for academics and industry clients Australia-wide and internationally with a focus on developing and executing high-throughput screening assays for biomarker and therapeutic discovery. More information about the ACTD facility and capabilities can be found here.







