Functional Genomics & High-Throughput Screening
Phenomics Australia enables researchers to perform genome-scale cell-based CRISPR, RNAi, and compound screening in both 2D cell lines and 3D cell lines, PDX, patient-derived cells, and complex disease models using sophisticated liquid handling automation, high content cellular phenotyping, and reporter-based readouts.
Expertise

-
CRISPR/Cas9 screening
-
Pooled viral CRISPR screening
-
Arrayed CRISPR screening
-
Nucleofection
-
CROPSeq
-
RNAi screening
-
Compound Screening
-
3D biology and screening
-
shRNA platform
-
Cellular phenotyping
-
Bioinformatics pipelines
-
Quantitative analysis
-
Data provision
Explainer
Functional genomics is the analysis of how individual (or multiple) genes contribute to a particular phenotype. Through functional genomics analyses, we are able to investigate the functional impact of newly discovered disease genes or variants identified in patients with rare diseases, allowing the development of new diagnostic and therapeutic outcomes.
High-throughput screening combines automated, robotic and liquid handling equipment with various sensing techniques to acquire assay data quickly and at scale. This capacity allows rapid functional interrogation of the genome, as well as the large scale screening of antibodies and drug libraries.
Phenomics Australia offers functional genomics and screening services for both cells and printed cells in scaffolds as physiological models.
Services

The CRISPR/Cas9 system enables gene deletion directly at the genomic level. This approach uses the bacterially derived enzyme Cas9 and a gene-specific single guide RNA molecule (sgRNA) to introduce nicks in precisely defined genomic locations leading to a complete gene knockout. This can be supported at the single target level, small scale boutique libraries or large whole genome campaigns.
Phenomics Australia houses the latest generation of genome-wide pooled CRISPR libraries to knockout protein coding genes (human Brunello and mouse Brie libraries), over-expression using CRISPRa (human Calabrese and mouse Caprano libraries) and inhibition using CRISPRi (human Dolcetto and mouse Dolomiti libraries). To identify genes responsible for the phenotype, we use next-generation sequencing and extensive bioinformatics analysis. These pipelines are available in conjunction with the facilities providing the screening resources.
Phenomics Australia supports both viral and synthetic arrayed CRISPR screening. Phenomics Australia has the TransEDIT dual guide viral library in arrayed format targeting every gene in the human genome, with 2 guides per gene per construct. This can be used for single target optimisation or validation of pooled screens, or as a boutique pooled library screening approach. The synthetic CRISPR human Horizon Discovery sgRNA Edit-R library enables gene knockout in an arrayed format using lipid or nucleofection delivery strategies in 96 or 384 well format. This platform uses synthetic purified sgRNAs that are transiently delivered into the Cas9-stable cells and the phenotype is assessed typically within 72-96h post-transfection. This platform enables cell phenotyping approaches using high content imaging.
Phenomics Australia houses the NucleofectorTM Technology that enables highly efficient transfection of primary cells, stem cells, neurons, and cell lines via electroporation. It is a commonly used method to deliver Cas9/sgRNAs ribonucleoproteins (RNPs) to your target cells.
CROPSeq experiments combines genetic perturbations with single cell RNA seq (scRNA-Seq). The result is a complete profiling of the total mRNA expression changes in a particular cell following CRISPR mediated deletion (CRISPRc) or suppression (CRISPRi) of 60-100 genes. The CROPSeq service includes all of the experimental and computational requirements for a CROPSeq experiment.
The siRNA platform, based on siGENOME SMARTpool reagents (Horizon Discovery) can be screened in arrayed formant in either 96 or 384 well plate format. siRNAs targeting the protein coding genome, miRNAs (mimic or inhibitor) or long non-coding RNAs are transiently introduced into cells using lipid-based reagents and assayed between three to seven days post-transfection. Screeners can perform imaging based high throughput, high content cellular analysis or fluorescence/luminescence based phenotypic quantitation using a high throughput plate reader. Often, researchers multiplex their screens and use both platforms.
Using high throughput liquid handling automation, Phenomics Australia enables all types of compound screening approaches from cell-based screens to complex, cell-free protein interaction screens. From straightforward dose curves with BYO drugs and multiple cell lines, to large compound collections, there is great creativity in this screening approach and we work with you to develop the assay readouts. Compounds are primarily sourced from Compounds Australia (Griffith University).
Using liquid handling automation or Rastrum technology, we can embed cells in scaffolds such as Matrigel or customised hydrogels in high throughput and can vary composition and stiffness. This enables exploration of cell lines, patient tissues and iPSCs to derive a picture of growth kinetics, response to targeted gene engineering and drug discovery. This platform enables highly complex co-culture interaction studies, including tumour and immune cells, host-pathogen interactions, angiogenesis and cell invasion. It has great potential for the new era of personalised medicine.
In situations where gene deletion is considered less desirable, Phenomics Australia has an arrayed lentiviral shRNA-mir30 vector system (pGIPZ) library that enables stable knockdown in primary, non-dividing and standard cell lines. shRNAs can be screened singularly for validation purposes or the whole genome screened as 10 pools. The facility with assist with our next-generation sequencing pipeline to identify the genes responsible for the biological phenotype of an assay.
Phenomics Australia screening facilities are underpinned by sophisticated high content microscopy platforms that enable researchers the capacity to quantify cellular phenotypes in response to perturbation (e.g. CRISPR, RNAi, compounds, 2D or 3D) in an entirely unbiased manner. High content microscopy is a fully automated method of imaging cells stained with relevant biological agents, simultaneously measuring thousands of cellular features. Using complex bioinformatics pipelines, we can identify cellular features that are significantly enriched in populations of cells treated with the same agents, allowing us to cluster like outcomes and triage our hit lists rapidly. This information can be overlaid with transcriptional and proteomic profiling data alongside available patient clinical information.
Big data requires innovative ideas and a lot of storage and analysis capabilities. Our screening facilities have data analysts who are expert cell biologists who understand the biological questions and implications of the screen outcomes and can run the software and derive customised analysis pipelines. They are a fundamental part of the success of every screen project. It is also essential to be in discussion about data analysis considerations at the start of each project to ensure the best possible experimental design is implemented.
The majority of screens focus on understanding cellular changes with an increasing focus on quantitative high content imaging. Common screening approaches being utilised are directed toward understanding drug resistance or drug sensitivity mechanisms, or identification of synthetic lethal targets that augment a cell death response in the combination of the gene and drug.
All data generated by Phenomics Australia facilities is owned by the research team running the project. The facilities will store and manage your data and analysis in accordance with the funding body requirements. Large data deposits generated through imaging will be expected to be taken by the research team at the conclusion of the project.
Further Information
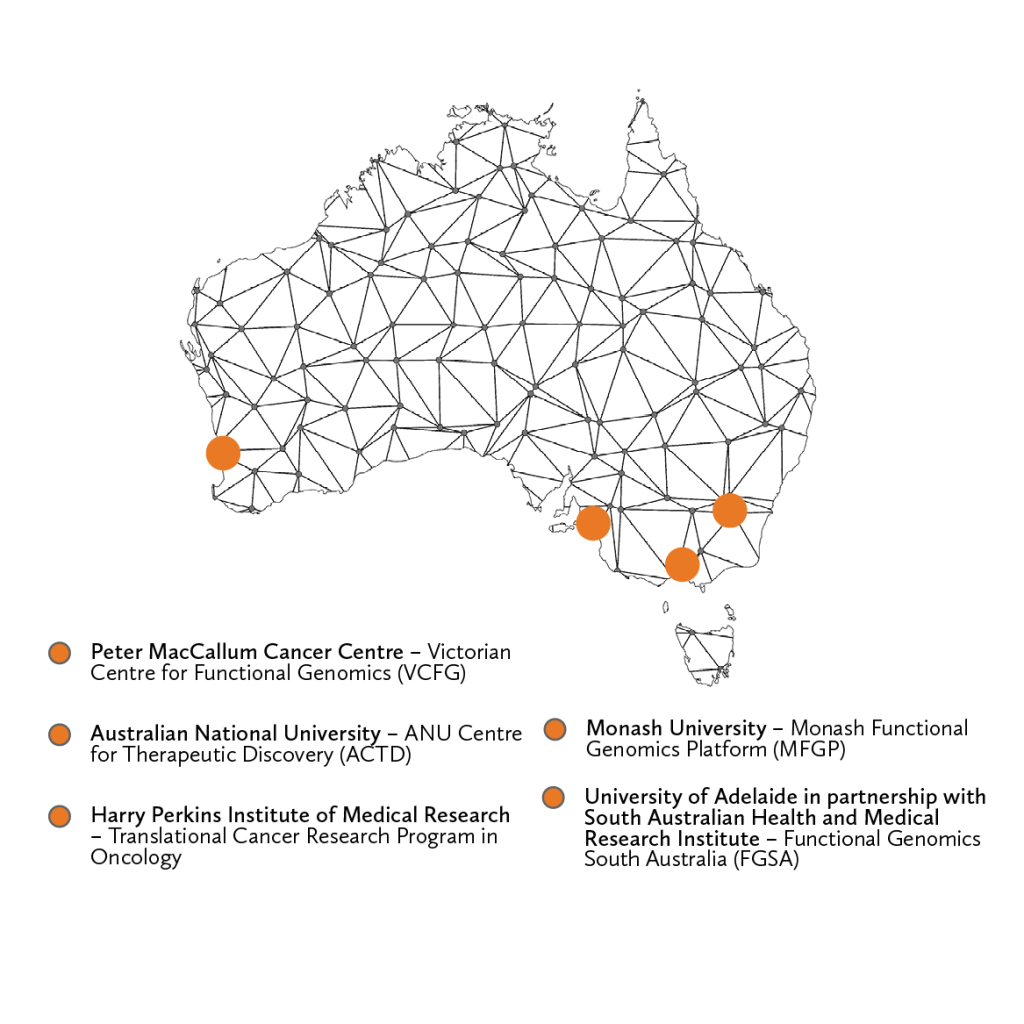
Through the Victorian Centre for Functional Genomics (VCFG) at the Peter MacCallum Cancer Centre, the Harry Perkins Institute of Medical Research (Perkins), the ANU Centre for Therapeutic Discovery (ACTD, The John Curtin School of Medical Research, ANU) the Monash Functional Genomics Platform (MFGP) at Monash University, and the Functional Genomics South Australia (FGSA) at the University of Adelaide (in partnership with SAHMRI), Phenomics Australia Functional Genomics and High-throughput screening services enable biomedical researchers Australia-wide with the ability to perform novel discovery-based screens using multiple platforms.
The facilities are staffed by experts and offer a range of access opportunities, from enabling researchers to be embedded in the facility and to be trained on appropriate equipment and fully supported by the team from assay concept design, screening and analysis, through to full fee-for-service screening capacity. Through this collaborative approach, the Researcher always maintains control of their data through frequent discussion with the facility team.
By housing extensive liquid handling automation, high content imaging capabilities and sophisticated plate reader instruments, the scale of a screen is entirely up to the researcher. All sites house an arrayed whole genome human synthetic CRISPR library and have access through the VCFG to arrayed and pooled viral library screening resources. The ACTD and VCFG also house whole human genome siRNA SMARTpool arrayed libraries for RNAi screening. The VCFG also houses human miRNA, long non-coding RNA and shRNA screening libraries and a mouse whole genome RNAi library. Through Compounds Australia, who house Australia’s largest compound repository at Griffith University, all sites are able to offer compound screens at any scale.
More information available on Phenomics Australia Resources page.